Oligomerization of rhodopsin and other G-protein-coupled receptors (GPCRs)
G-protein-coupled receptors (GPCRs) constitute a large superfamily of receptor proteins responsible for signal transduction (about 800 types). These receptors mediate recognition of environmental stimuli like light, odor, and taste, but also involve responses to peptides, hormones, proteases, chemokines and other ligands across plasma membranes. They are also important targets for pharmacological intervention via activating or blocking their action. The current state of research is that GPCRs exist and act as dimers. However, a couple of years ago the monomeric state of GPCR was commonly accepted and a few cases of dimerization (like GABAB receptors) were treated as an exception. Gradually, due to growing evidence of experimental data the hypothesis of dimerization became a dominant one.
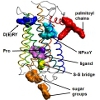
Based on distances between rhodopsin monomers, measured by Atomic Force Microscopy (AFM), we built a model of rhodopsin oligomer. The model was subsequently enhanced by simulation in membranes specific to native rod outer segment (ROS) discs. Such membranes contain unsaturated lipids (docosahexaenoyl chains) in high concentrations. Our model of rhodopsin oligomer (1N3M in Protein Data Base) was experimentally confirmed not only for rhodopsin but also for dopamine receptor. It can suggest that functional dimers of GPCRs may employ similar interfaces as well as mechanisms of signal transduction.
- July 2023.GS-SMD web server
for SMD simulations of γ-secretase complex.
Publication in Nucleic Acids Research 2023 Web server issue.
- August 2022.COGRIMEN
- June 2021.GPCRsignalOur new service GPCRsignal was recently published in NAR 2021, W1.