Alzheimer's Disease - structure of gamma-secretase complex
Presenilin-1 (PS-1) is a catalytic component of the membranous complex of γ-secretase which plays an important role in onset and development of Alzheimer’s Disease. It cuts the APP (Amyloid Precursor Protein) to β-amyloid which can aggregate in neurons. The γ-secretase complex consists of four proteins: PS1, APH-1, PEN-2 and nicastrin (Nct). Structures of all of these proteins are unknown. In the maturation process the structure of PS-1 is cleaved on N- and C-terminal fragments (NTF and CTF). Structure of CTF was determined using NMR methods in detergent micelles (Sobhanifar et al., PNAS 2010). Our group revealed how this structure is changing in a micelle of detergent molecules as well as in lipid bilayers (DLPC, DPPC and implicit) using different types of molecular dynamics simulations. This research is continued toward determination of interactions with the substrate APP and drug design.
(Cooperation with BioNMR Centre at Goethe University, Frankfurt/Main, Germany)
- July 2023.GS-SMD web server
for SMD simulations of γ-secretase complex.
Publication in Nucleic Acids Research 2023 Web server issue.
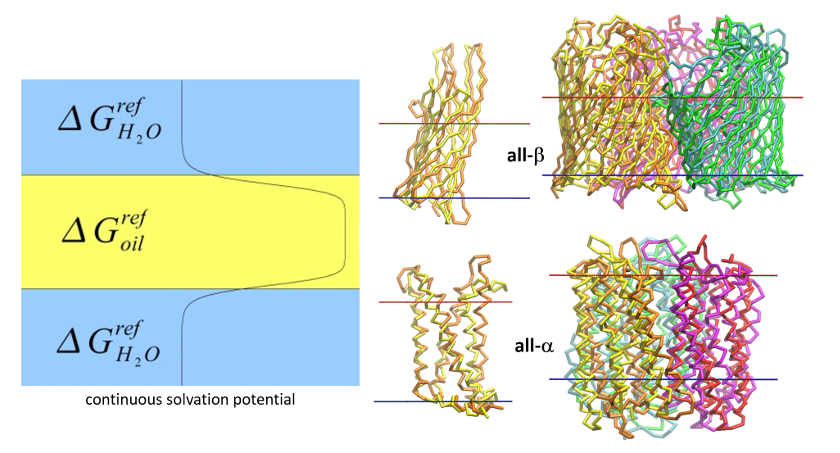
- August 2022.COGRIMEN
- June 2021.GPCRsignalOur new service GPCRsignal was recently published in NAR 2021, W1.